Recently, I have developed a tool named MetaLogo, aimed to make sequence logos for multiple sets of sequences.
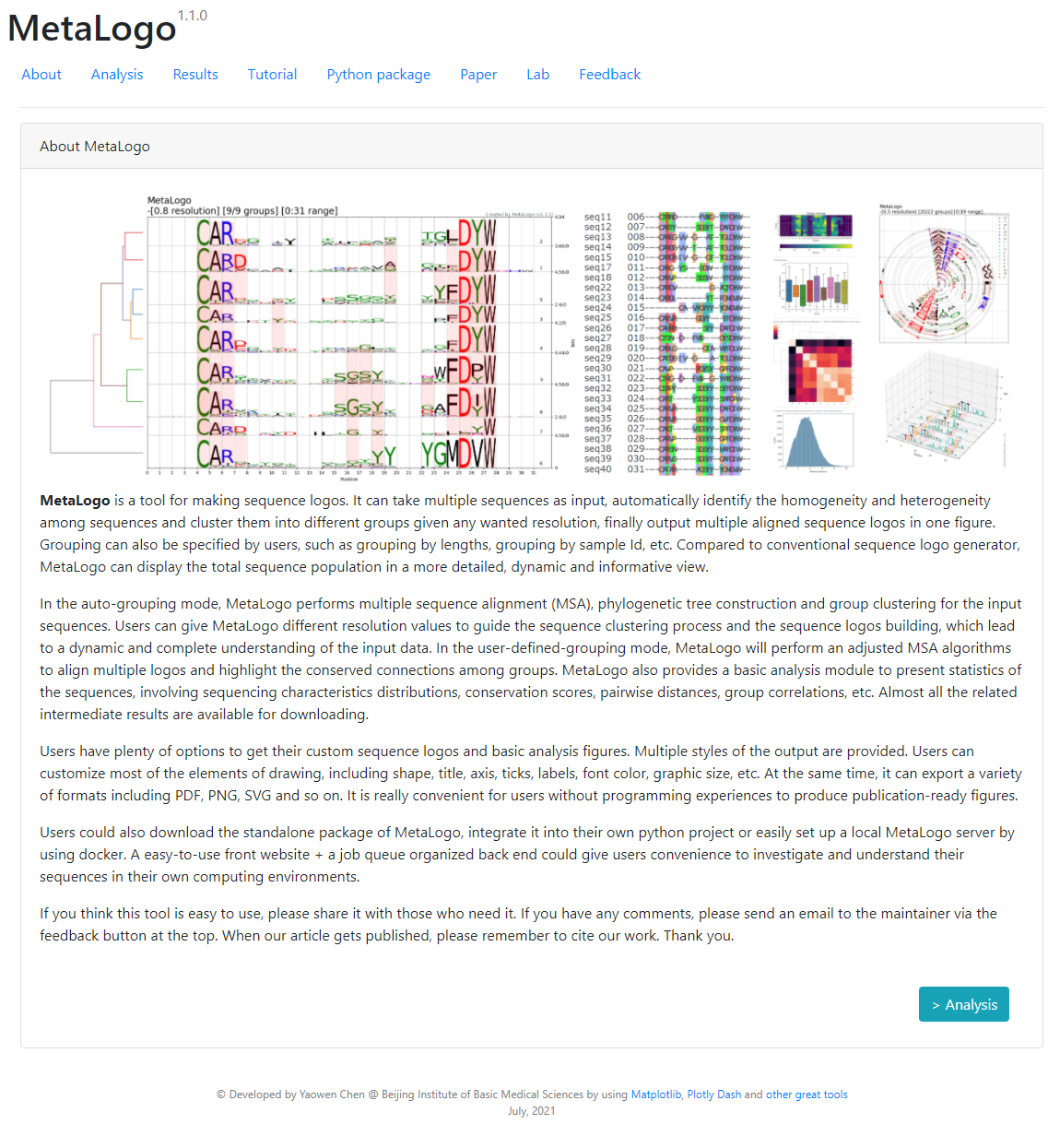
MetaLogo is a tool for making sequence logos. It can take multiple sequences as input, automatically identify the homogeneity and heterogeneity among sequences and cluster them into different groups given any wanted resolution, finally output multiple aligned sequence logos in one figure. Grouping can also be specified by users, such as grouping by lengths, grouping by sample Id, etc. Compared to conventional sequence logo generator, MetaLogo can display the total sequence population in a more detailed, dynamic and informative view.
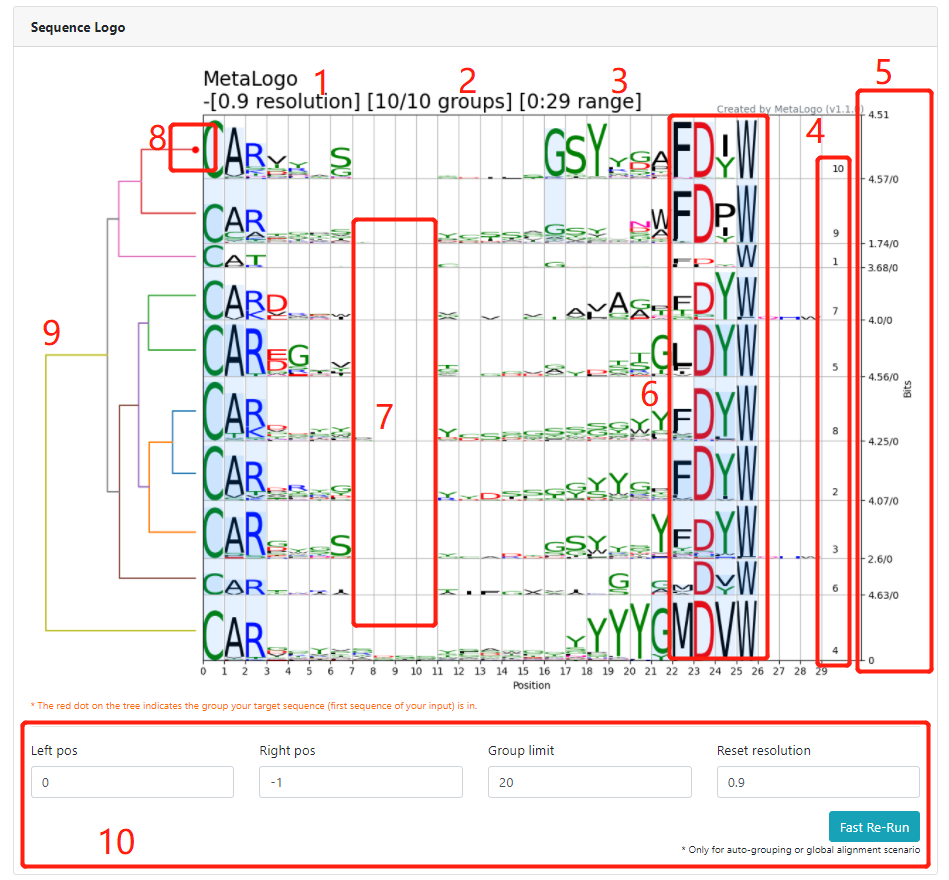
In the auto-grouping mode, MetaLogo performs multiple sequence alignment (MSA), phylogenetic tree construction and group clustering for the input sequences. Users can give MetaLogo different resolution values to guide the sequence clustering process and the sequence logos building, which lead to a dynamic and complete understanding of the input data. In the user-defined-grouping mode, MetaLogo will perform an adjusted MSA algorithms to align multiple logos and highlight the conserved connections among groups. MetaLogo also provides a basic analysis module to present statistics of the sequences, involving sequencing characteristics distributions, conservation scores, pairwise distances, group correlations, etc. Almost all the related intermediate results are available for downloading.
Users have plenty of options to get their custom sequence logos and basic analysis figures. Multiple styles of the output are provided. Users can customize most of the elements of drawing, including shape, title, axis, ticks, labels, font color, graphic size, etc. At the same time, it can export a variety of formats including PDF, PNG, SVG and so on. It is really convenient for users without programming experiences to produce publication-ready figures.
Users could also download the standalone package of MetaLogo, integrate it into their own python project or easily set up a local MetaLogo server by using docker. A easy-to-use front website + a job queue organized back end could give users convenience to investigate and understand their sequences in their own computing environments.
继续阅读Make sequence logo for heterogeneous sequences or sequences with unequal length using MetaLogo?